Background
The crux of sexual reproduction in multicellular organisms is that all information required for the development of that organism is present entirely in the sperm and the oocyte. The vast majority of this information is present in the DNA, which carries all the genes that encode the proteins required for cellular function and viability. However, beyond the DNA blueprint provided by the gametes, other factors delivered during fertilization are required to initiate embryogenesis, and to orchestrate a robust cascade of differentiation events required for multicellular development.
Many maternally supplied factors have been demonstrated to be essential for development. As one prominent example, RNAs and proteins present in the oocyte are required to initiate the first cellular division after fertilization, and for the consequent activation of transcription of the zygotic genome. Due to the discrepancy in size of sperm and oocyte (100 to 1000-fold in most organisms) it has been thought that sperm contribute little or nothing to early development beyond their haploid genome and in some organisms such as humans the first functional centriole.
More recently, however, it has become increasingly understood that sperm transmit molecules including RNAs during fertilization thereby eliciting non-genetically inherited phenotypes in offspring. This understanding has come predominately through the study of paternal environmental effects, where paternal environmental exposures or experiences alter the phenotype of succeeding generations. Small (regulatory) RNAs such as microRNAs (miRNAs) and tRNA-Fragments (tRFs) have been identified as a carrier of environmentally-modulated sperm-transmited non-genetic information in organisms ranging from C. elegans (worms) to mice.
Currently, the mechanisms explaining how RNA transmitted by sperm to the zygote during fertilization can modulate the phenotype of offspring are completely unknown.
The Conine Lab is interested in how non-genetic inheritance occurs at a molecular level. What sperm RNAs are able to regulate early embryonic development and transmit information to offspring? How do sperm RNAs regulate embryonic gene expression and development - or what are the first alternations to development that sperm RNAs elicit? Finally, how do sperm RNAs alter the cascade of developmental events necessary to to produce an organism, thereby modulating offspring phenotype?
To answer these questions, the lab uses various model systems, including mice and worms to produce a combined genetic, molecular, biochemical, and genomic framework for determining the mechanisms underlying the functions of sperm-transmitted RNAs.
Samantha
May
Marketing Executive
I'm a paragraph. Click here to add your own text and edit me. It’s easy. Just click “Edit Text” or double click me.
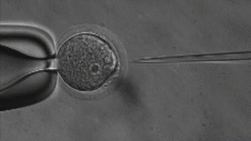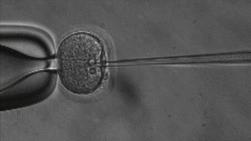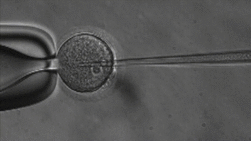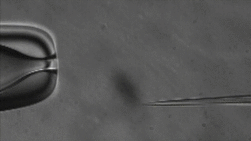
ICSI
The lab uses Intracytoplasmic Sperm Injection (ICSI); This Assisted Reproduction Technique (ART) allows for fertilization of an egg via injection of an individual sperm, producing a zygote.